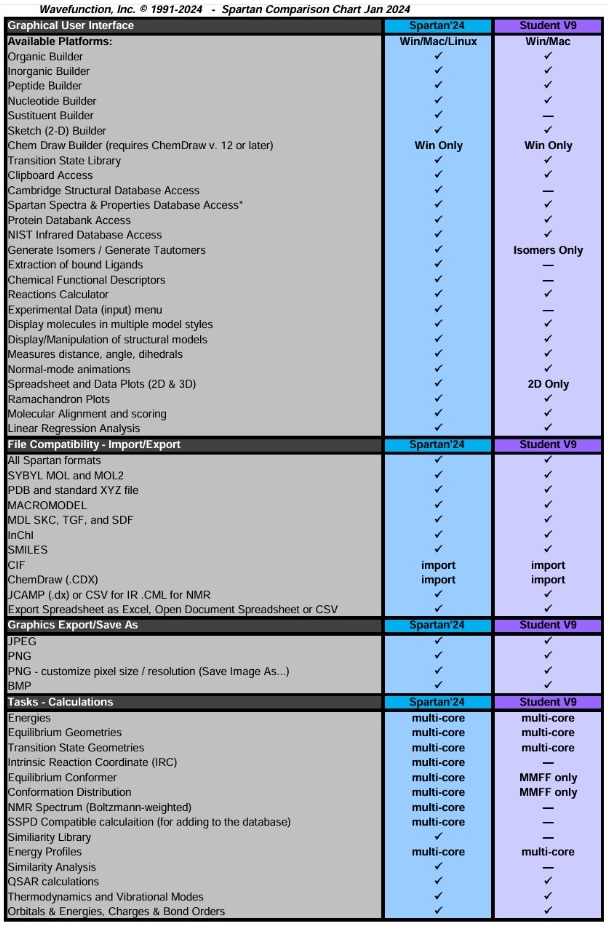
最新版 Spartan'24 Parallel Suite 更新於 2024/1/1
最新版 ODYSSEY v7
Spartan'24 for Win, Mac, & Linux
Spartan是目前提供版本裡最先進成熟的。最新版本提供所有的功能和方法,這些都包括在Spartan Essential版,並且與Q-Chem合作,提供了一個全方位的post-Hartree-Fock方法,包括密度功能、Moller Plesset,Theremochemistry訣竅(包括新的T1程序),以及相關類別的先進做法。 用
工業界和學術界化學研究的終極桌面應用程式的最新版本。透過業界最直觀的使用者介面可以獲得全方位的理論模型。比以往任何時候都更增強、精緻和更快。Spartan'24許可證配置為利用最多 16 個核心來執行選定任務以及Spartan Spectra 和屬性資料庫,並且能夠充當從其他Spartan許可證或iSpartan應用程式(在 iPad、 iPhone 或 iPod Touch)。
前所未有的化學計算
Spartan是最新發布了波函數旗艦Spartan線。除了性能,穩定性以及提供了超過17年的專業軟體開發功能,以下更添加了新的功能。
點選圖片可放大觀賞
| Equilibrium Geometry Structures | .jpg) |
Molecular Properties | .jpg) |
| Calculated Electronic Surfaces | .jpg) |
Conformational Analysis | .jpg) |
| 3D Alignment & Superposition | .jpg) |
3D Similarity Analysis | .jpg) |
Spartan Student Edition for Windows/Macintosh
終於,一個可以給教學或學習用的化學軟體誕生了,它是一個非常重要的分子建模套件,結合Spartan的易於使用圖形界面,主要的功能是針對計算方面,以及在化學教育裡提供了前所未有的分子模擬技術的波函數
Spartan Student Edition provides affordable molecular modeling software that takes advantage of modern computational methods to explore fundamental concepts in general, organic, physical, and inorganic chemistry courses. The graphical interface is based on the latest Spartan release. The computational features are a targeted subset of those in the full Spartan version. Click HERE for a feature comparison. Spartan is used in chemistry education and research at thousands of colleges, universities, government agencies, and commercial sites, worldwide.
Please note: Licensing is intended solely for student purchase for use on Student-owned machines. Licensing is machine locked and cannot be transferred. For installation on campus machines, please visit the Academic License Purchase Page. Wavefunction may require proof of student status before releasing activation codes. Please use your student email address when ordering.
Automated Conformational Analysis
A multi-step recipe for obtaining accurate Boltzmann distributions has been implemented. Combining MMFF, Hartree-Fock, and density functional models, Boltzmann distributions are ultimately obtained from either B97M-V/6-311+G(2df,2p)[6-311G*] energies with B3LYP/
6-31G* geometries or wB97X-V/6-311+G(2df,2p)[6-311G*] energies with wB97X-D/6-31G* geometries. User-customization is available for theoretical models used to obtain geometry and energy.
Automated Nmr For Flexible Molecules
Automated conformational analysis along with NMR chemical shift
calculations from empirically corrected B3LYP/6-31G* or wB97X-D/6-31G*
models provide Boltzmann-averaged NMR spectra for flexible molecules.
Dp4 Support
The DP4 measure is available to determine the best match of calculated and experimental
NMR chemical shifts from among a selection of stereo/ regio isomers (or the Boltzmann
averaged NMR of each isomer) using either B3LYP/6-31G* orwB97X-D/6-31G* models.
Coupling Constants
Two, three, four and higher-bond HH, CH and CC coupling constants may be calculated in full or by using only the Fermi contact term. CH coupling constants allow calculation of realistic 2D HMBC plots. For backward compatibility, three-bond HH coupling constants from an extended form of the Karplus equation remain available.
2d Sketches From 3d Structures
3D to 2D structure conversion is available and resulting 2D sketches may
be copied/pasted into spectra and plots.
Modern Computational Methods
Spartan’18 Parallel Suite provides convenient access to a number of
important computational methods, with emphasis on density functional
models among them:
GGA functionals. B86PW91, BLYP,
BPW91, B97-D2, SOGGA11, PBE-D3,VV10
GH-GGA functionals. B3LYP,B3LYP-D3, EDF2, B3PW91, B97-3,MPW3LYP, SOGGA11-X
RSH-GGA functionals. wB97X-D,wB97X-V, wB97X, CAM-B3LYP, N12-SX,
LC-VV10
mGGA functionals. B97M-V, M06-L,BMK, M11-L, TPSS-D3
GH-mGGA functionals. M06-2X, M06, M08-HX, M08-SO, MPW1B95
RSH-mGGA functionals. M11, wB97M-V, MN12-SX
Among the wave function based or correlated models are QCISD,QCISD(T), CCSD and CCSD(T) as well as the electric component of G3,G3(MP2), G4 and G4(MP2).
Available basis sets include the full range of Pople sets, the widely-used
Dunning correlation consistent sets, and Ahlrich/Weigend def2 sets.
Triple-ζ basis sets are now available for transition metals and lanthanides,allowing improved reaction energy calculations. The dual basis set approximation may be employed with double-ζ and larger basis sets,leading to an order of magnitude reduction in computation time.
From menus:
Pople: STO-3G, 3-21G, 6-31G*, 6-31G**, 6-31+G**, 6-311G**, 6-311+G**,
6-311G(2d,p), 6-311+G(2d,p), 6-311+G(2df,2p), 6-311+G(3df,2p);additional variations on the underlying 6-31G and 6-311G basis sets may be specified
Dunning: cc-pVDZ, aug-cc-pVDZ, cc-pVTZ, aug-cc-pVTZ, cc-pVQZ,aug-cc-pVQZ
Ahlrichs/Weigend: def2-SV(P), def2-SVPD, def2-TZVP, def2-TZVPPD,def2-QZVP, def2-QZVPPD
Remote Submission
Spartan’18 Parallel Suite can be used as a server for both remotely
submitted calculations and the Spartan Spectra and Properties Database,extending access to computational tools from remote devices. Remote computation is available from computers running Spartan’14 or later as well as iOS devices (iPad and iPhone) running the iSpartan app.
Graphical Interface
Sketch organic, inorganic, organometallic molecules in 2D and automatically convert to 3D structures. Groups, rings and ligands templates are available Build organic, inorganic and organometallic molecules, peptides and
nucleotides, substituted molecules in 3D Auto-convert 3D structures to 2D sketches
Link seamlessly to ChemDraw®Display/query molecules in a variety of model styles Display dipole vector, hydrogen bonds, points and planes. Display and customize chemical function descriptors Display user-defined annotations .Align molecules using structure, chemical function descriptors, or atom labels ,Align molecules to pharmacophores. Build transition states using reaction arrows . Generate transition states from an extensive reaction library . Generate and display molecular orbitals, electron densities, spin densities
electrostatic potentials, electrostatic potential maps, orbital maps, and local ionization potential maps
Optionally silhouette mesh and transparent surfaces for improved visualization
Generate and display orbital energy diagrams on-the-fly
Optionally view property maps in Red-White-Blue color scale
Toggle between default and absolute property ranges for property maps
Display electron density based on % of enclosed electrons
Highlight solvent accessible regions on surfaces and property maps
Display R/S chirality markers, invert chiral centers and absolute chirality
Integrated reaction energy calculator
Organize data in spreadsheets
Print and save calculated spectra tables as PDF files
Perform regression analysis and make, save and print 2D or 3D plots
Embed external files such as MS® Office and Adobe® PDF files in Spartan files
Plot, print, and save calculated and experimental IR, Raman, and UV/vis spectra
Display calculated 1D (proton,13C, DEPT) and 2D (COSY, HSQC, HMBC) and
experimental 1D proton and 13C NMR spectra
Overlay experimental HMBC and COSY 2D NMR spectra with calculated results
Tasks
Calculate strain energy, total energy and heat of formation
Determine gas-phase equilibrium and transition-state geometries
Determine geometries and IR spectra in the presence of solvent
Identify global minimum; calculate Boltzmann weights to obtain
conformer energy distributions
Build libraries of diverse conformers for use in similarity analysis
Perform similarity analysis on the basis of structure or chemical functionality
Scan geometrical coordinates and generate reaction sequences
Calculate reaction and activation energies
Calculate IR, Raman, UV/vis, and NMR spectra
Determine NMR spectra for flexible molecules
Match calculated and experimental NMR spectra
Search SSPD and NIST experimental database for match to calculated IR spectra
Mine databases of calculated molecular, atomic, and reaction properties
Computational Methods
Molecular Mechanics. SYBYL, MMFF94, MMFF(aq)
Semi-Empirical. MNDO, AM1, RM1, PM3 (with transition metal parameters), PM6
Hartree-Fock molecular orbital theory
Density Functional Theory (functionals available from menus):
GGA functionals: B86PW91, BLYP, BPW91, B97-D2, SOGGA11, PBE-D3, VV10
GH-GGA functionals: B3LYP, B3LYP-D3, EDF2, B3PW91, B97-3, SOGGA11-X
RSH-GGA functionals: ωB97X-D, ωB97X-V, ωB97X, CAM-B3LYP, N12-SX, LC-VV10
mGGA functionals: B97M-V, M06-L, BMK, M11-L, TPSS-D3
GH-mGGA functionals: M06-2X, M06, M08-HX, M08-SO, MPW1B95
RSH-mGGA functionals: M11, ωB97M-V, MN12-SX
Functionals may be customized and additional functionals specified via keywords
Møller Plesset. MP2, MP3, MP4, and RI-MP2
Wave function based advanced correlated models:
G3(MP2)elect, G3elect, G4(MP2)elect, G4elect, QCISD, QCISD(T), CCSD, CCSD(T)
Additional wave function based advanced correlated methods are available
from keywords and include CCSD, CCSD(T), OD, OD(T), QCCD, VOD, and VQCCD
Excited-state methods. CIS, CIS(D), RI-CIS(D), QCIS(D), QCISD(T) and TDDFT
Gradients available for CIS, CIS(D) and TDDFT
Thermochemical Recipes. T1, G3(MP2), G3, G4(MP2), G4
Basis Sets (available from menus):
Pople: STO-3G, 3-21G, 6-31G*, 6-31G**, 6-31+G**, 6-311G**, 6-311+G**,
6-311G(2d,p), 6-311+G(2d,p), 6-311+G(2df,2p), 6-311+G(3df,2p); other variations
on 6-31G and 6-311G may be specified
Dunning: cc-pVDZ, aug-cc-pVDZ, cc-pVTZ, aug-cc-pVTZ, cc-pVQZ, aug-cc-pVQZ
Ahlrichs/Weigend: def2-SV(P), def2-SVPD, def2-TZVP, def2-TZVPPD, def2-QZVP,
def2-QZVPPD
Automatic use of pseudopotentials for elements >Kr (including lanthanides and
select actinides)
Dual basis set approximation available with double-z and larger basis sets
Import custom basis sets
Properties And Qsar Descriptors
Mulliken, natural, and electrostatic-fit charges
Dipole and higher moments, polarizabilities and hyperpolarizabilities
Enthalpies, entropies and free energies
Solvation energies from C-PCM, as well as SM5.4, SM8, SM12, SMD and SS(V)PE
Statistical tools for comparing calculated and experimental 13C NMR shifts
HOMO, LUMO and SOMO energies
Areas, polar surface areas and volumes based on space-filling models
Areas, accessible areas, polar areas, and volumes based on the electron density
Min/Max of electrostatic potential and Min of local ionization potential
Number of conformers and tautomers
Number of hydrogen bond acceptors and donors
Spectra
NMR, IR, Raman and UV/visible spectra may be calculated using a variety of
theoretical models: Hartree-Fock and density functional models for NMR, semiempirical, Hartree-Fock, density functional and MP2 models for IR, Hartree-Fock,density functional models for Raman and UV/visible.
NMR
A 3rd-generation parameterization scheme for corrections to NMR chemical shifts
for the B3LYP/6-31G*, wB97X-D/6-31G* and wB97X-D/6-311G* models is available.
Calculated (corrected) chemical shifts together with calculated three-bond HH,
CH, and CC coupling constants (using B3LYP or wB97X-D functionals with PCJ-0,
PCJ-1, or PCJ-2 basis sets) allow for a 1
H 13C and DEPT 1D spectra and COSY,
HSQC and HMBC 2D spectra. Proton spectra may be displayed with or without
HH coupling.
IR and raman
Infrared (and Raman) frequencies calculated from ωB97X-D/6-31G*, B3LYP/6-31G*
and EDF2/6-31G* density functional models are scaled to account for systematic
errors associated with the harmonic approximation. Corrected frequencies and
intensities are fit to a Lorentzian function with a line width parameter.
Alternatively, scale and line width may be adjusted to best fit a spectrum calculated using
any theoretical model to an experimental spectrum.
uv/visible
UV/visible spectra are obtained by explicit calculation of the ground state
energies and the low-lying excited states. CIS models are paired with HartreeFock models. TDDFT (time dependent density functional) models are paired with density functional models.
Additional Features
Multi-core parallel processing for Hartree-Fock, density functional, RI-MP2, and
thermochemical recipes.
Automatic processing of groups of molecules
Automatic use of molecular symmetry
View recent documents from File menu
Optimize using constraints and/or frozen atoms
NOEs for conformational searching
Identify tautomers and generate tautomer lists
Import experimental IR, Raman, and NMR spectra
Import structures in InChI, SMILES, CDX, CIF, SKC, SDF, TGF, XYZ, Macromodel, PDB,
SYBYL MOL and MOL2 format
Retrieve structures from Cambridge Structural Database and Protein Data Bank
Extract ligands and binding sites from proteins (PDB files)
Databases
Spartan Spectra and Properties Database (SSPD)® comprises two collections, the
first of ≈300,000 organic molecules obtained from the EDF2/6-31G* model and
the second of ≈300,000 organic molecules and ≈2,000 organometallic molecules
obtained from the wB97X-D/6-31G* model. Both include the optimized geometry,the energy and a selection of molecular properties, the wave function (allowing on-the-fly generation of graphical surfaces), and the NMR spectrum. The infrared spectrum is also provided for molecules in the EDF2/6-31G* collection. Individual SSPD entries can replace user-built structures, and both collections are searchable by substructure, name, formula, and isomer.
Spartan Reaction Database (SRD)® comprises transition states for ≈1,800 reactions
searchable by combination of substructure and “reaction arrows” from either 2D
sketches or 3D models
ωB97X-V/6-311+G(2df,2p) ENERGY DATABASE
SSPD entries now include calculated energies from the wB97X-V/6-311+G(2df,2p)
model, providing more accurate reaction energies than provided by the wB97X-D/6-31G* model. These can be accessed from the Properties or Reactions dialogue as well as from the Spreadsheet.
Enhanced Parallel Performance
Parallelization now includes shared-memory frequency calculations. Performance
has been improved for > 8 core systems.
Summary Output
Provides Tables for NMR chemical shifts and coupling constants, IR and Raman
frequencies and intensities and UV/visible absorption frequencies and strengths
are provided and may be saved as PDF files
Chemical Shift Labels
Labels for experimental NMR chemical shifts and differences between calculated
(Boltzmann averaged) and experimental chemical shifts are now available.
WINDOWS:
Modern Intel or AMD Processors (64-bit Only)
Windows 10 or 11
4 GB RAM
128 GB disk space or higher (SSD recommended)
MACINTOSH:
Intel or Mx Chip Only
4 GB RAM or higher
OS 10.12 (Sierra) through OS 14.X (Sonoma)
128 GB disk space or higher (SSD Recommended)
LINUX:
Modern Intel or AMD Processors (64 bit Only)
Linux RHEL 8, CentOS 8, Ubuntu 20.04 LTS
4 GB RAM
128 GB disk space or higher (SSD recommended)
  ODYSSEY®是一種獨特的教學方案,在一般高中、學院和大學中用來介紹基礎化學課程。利用科學為基礎的分子模擬,ODYSSEY提供了一個互動的環境,以便於學習和探索。 |
OD YSSEY 特別適合
YSSEY 特別適合
課堂報告
實驗室試驗
家庭作業
ODYSSEY Student Edition
這是一個化學學習的工具,它提供了一個管道讓您能夠直接進入分子的世界。充分發展的實驗過程,有一個大規模的樣品室和一個易於使用的模型套件,用來建築幾乎所有系統 – ODYSSEY提供給學生一個虛擬實驗室,用來學習基礎的發現力是非常理想的。也能讓您探索前所未有的化學!
ODYSSEY Instructor Edition
尖端教學軟體,可立即使用化學實驗和學生分配表。包括註明分子樣品室和一個易於使用的模型套件,可以用來建設幾乎所有的化學系統。對任何在分子水平的可視化化學感興趣的科學老師來說,這是一個新的必備工具。
ODYSSEY Release 7.0 新版介紹
• Succinct tutorials for the general operation of the program
• Expanded step-by-step instructions for building new models
• Learning Units for more than 290 topics
• Improved method for earmarking and accessing preferred labs
• More than 110 Google Forms for assessment purposes
• Wider adoption of terms commonly used in a teaching context
• Wider search functionality
• Molecular models for more than 800 compounds and mixtures
• Molecules/Ions as well as models of Gases/Liquids/Solids
• Learning units in Japanese (分子化学)
• Learning units in Spanish (Materia en movimiento)
• Learning units in German (Teilchen und Materie)
• Integration with Chemistry: Human Activity, Chemical Reactivity by Mahaffy, Tasker, Bucat, Kotz, Treichel, Weaver, and McMurry (Nelson Education)
• Extensively revised and expanded content (new units for lattice energy, -bonding, tautomers, and more)
• Improvements/corrections for dipole and ion visualization; charge labeling of all ions
• Compounds of higher-row elements shown with bond types that deemphasize octet expansion
• New models added in the Stockroom
• Answer key for more than 1,000 questions
• Misconceptions listed for major learning units
• Saving of complete molecular labs (can be hyperlinked/distributed)
• Hyperlinks to online teaching tutorials
• Alignments for secondary school usage (available from Help menu)
• Customization of worksheet questions
• Enable Authoring of New Labs (optional in Preferences)
系統需求
WINDOWS:
2 GHz chip or faster
Windows 8.1, 10, and 11
128 GB disk space or higher (SSD Recommended)
4 GB RAM
MACINTOSH:
Intel or M1 chip Macintosh (only), 2GHz chip or faster
OS 10.12 (Sierra) through OS 14.X (Sonoma)
128 GB disk space or higher (SSD Recommended)
4 GB RAM or higher